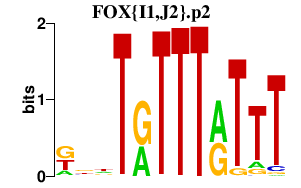
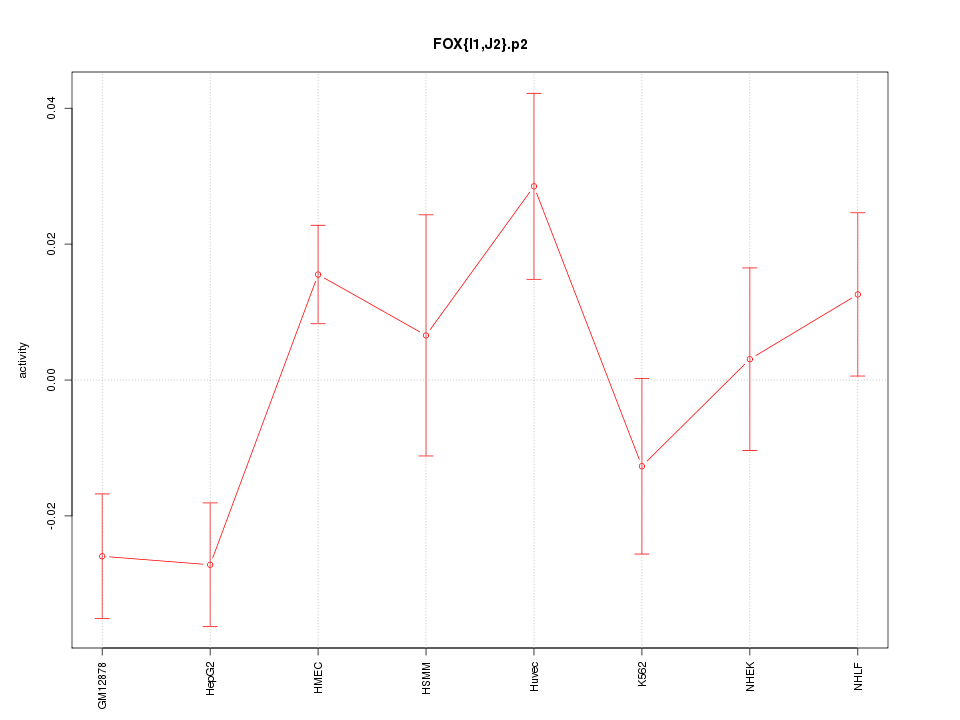
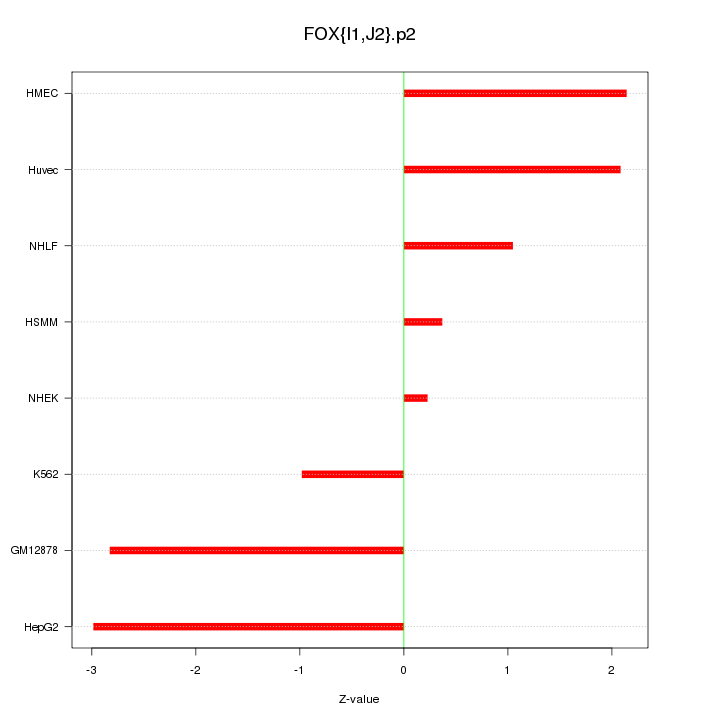
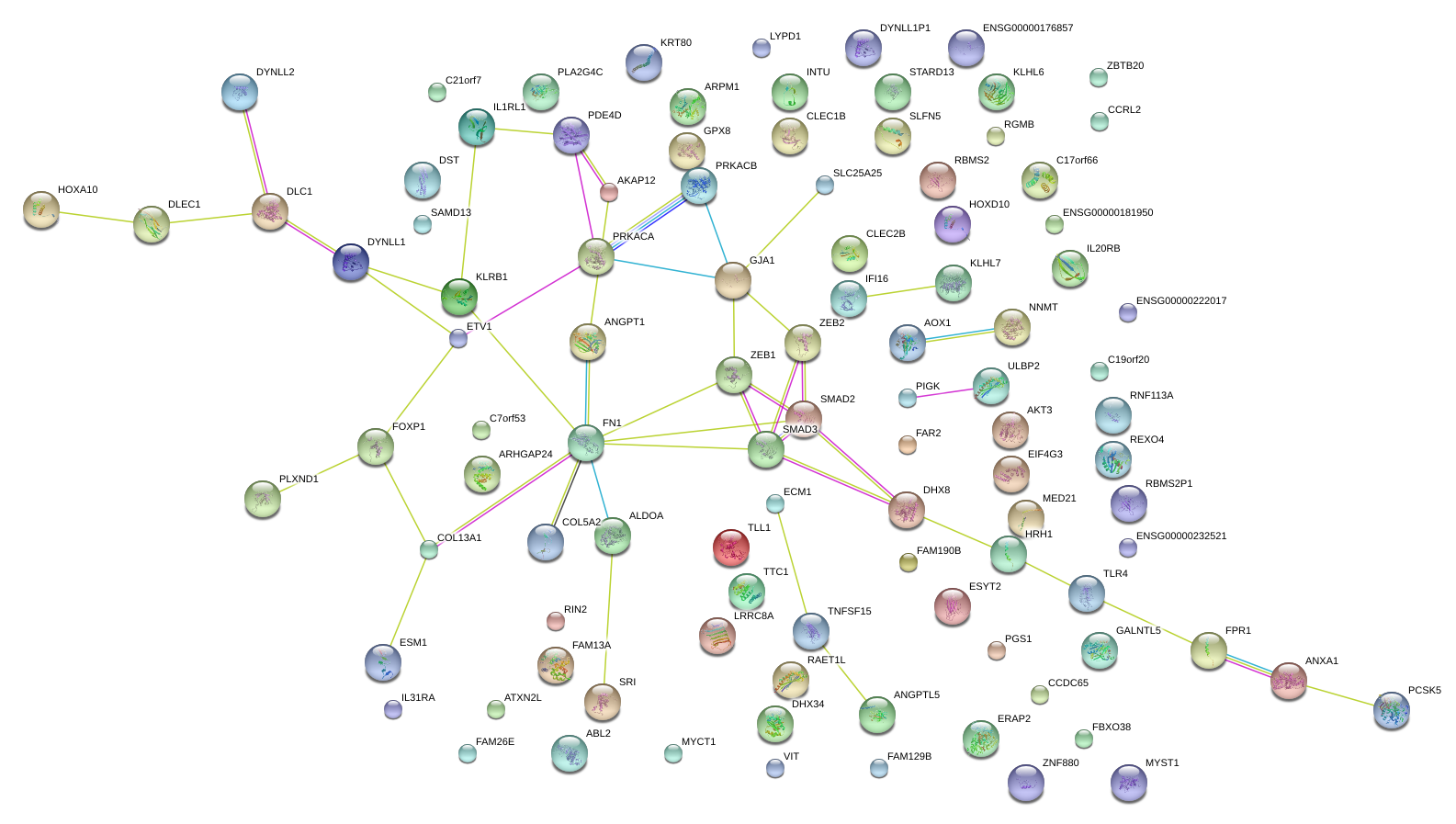
|
chr20_+_19818209
|
9.986
|
NM_018993
|
RIN2
|
Ras and Rab interactor 2
|
|
chr5_+_159368690
|
5.965
|
|
TTC1
|
tetratricopeptide repeat domain 1
|
|
chr5_+_159368733
|
5.225
|
NM_003314
|
TTC1
|
tetratricopeptide repeat domain 1
|
|
chr5_+_159368721
|
5.183
|
|
TTC1
|
tetratricopeptide repeat domain 1
|
|
chr4_+_167013859
|
4.112
|
NM_012464
|
TLL1
|
tolloid-like 1
|
|
chr5_-_59100194
|
3.642
|
NM_001197218
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr13_-_32757835
|
3.588
|
NM_178006
|
STARD13
|
StAR-related lipid transfer (START) domain containing 13
|
|
chr6_+_134800533
|
3.529
|
|
LOC154092
|
hypothetical LOC154092
|
|
chr3_+_11242666
|
3.446
|
NM_001098211
|
HRH1
|
histamine receptor H1
|
|
chr7_+_151284396
|
3.236
|
NM_145292
|
GALNTL5
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase-like 5
|
|
chr5_+_147775652
|
3.205
|
|
FBXO38
|
F-box protein 38
|
|
chr4_+_86968069
|
3.171
|
|
ARHGAP24
|
Rho GTPase activating protein 24
|
|
chr20_+_19863716
|
3.170
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr9_+_74956601
|
2.993
|
|
ANXA1
|
annexin A1
|
|
chr8_-_108579270
|
2.876
|
NM_001146
|
ANGPT1
|
angiopoietin 1
|
|
chr9_+_74956494
|
2.762
|
NM_000700
|
ANXA1
|
annexin A1
|
|
chr1_-_77457664
|
2.651
|
NM_005482
|
PIGK
|
phosphatidylinositol glycan anchor biosynthesis, class K
|
|
chr6_+_150304813
|
2.638
|
NM_025217
|
ULBP2
|
UL16 binding protein 2
|
|
chr5_-_54317102
|
2.630
|
NM_001135604
NM_007036
|
ESM1
|
endothelial cell-specific molecule 1
|
|
chr1_-_21373816
|
2.601
|
NM_001198803
|
EIF4G3
|
eukaryotic translation initiation factor 4 gamma, 3
|
|
chr2_+_201182187
|
2.564
|
|
AOX1
|
aldehyde oxidase 1
|
|
chr9_+_77695334
|
2.535
|
NM_001190482
NM_006200
|
PCSK5
|
proprotein convertase subtilisin/kexin type 5
|
|
chr6_+_153060722
|
2.534
|
NM_025107
|
MYCT1
|
myc target 1
|
|
chr7_+_111908143
|
2.492
|
NM_001134468
NM_182597
|
C7orf53
|
chromosome 7 open reading frame 53
|
|
chr21_+_29439768
|
2.425
|
|
C21orf7
|
chromosome 21 open reading frame 7
|
|
chr12_+_27066768
|
2.389
|
|
MED21
|
mediator complex subunit 21
|
|
chr11_+_113672378
|
2.350
|
|
NNMT
|
nicotinamide N-methyltransferase
|
|
chr15_+_65205107
|
2.344
|
NM_001145102
|
SMAD3
|
SMAD family member 3
|
|
chr9_+_129900637
|
2.341
|
NM_052901
NM_001006643
|
SLC25A25
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 25
|
|
chr3_-_170970296
|
2.337
|
NM_032487
|
ARPM1
|
actin related protein M1
|
|
chr3_+_46423682
|
2.337
|
NM_003965
NM_001130910
|
CCRL2
|
chemokine (C-C motif) receptor-like 2
|
|
chr19_+_57564981
|
2.326
|
NM_001145434
|
ZNF880
|
zinc finger protein 880
|
|
chr9_+_129900590
|
2.255
|
|
SLC25A25
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 25
|
|
chr11_+_113672407
|
2.238
|
|
NNMT
|
nicotinamide N-methyltransferase
|
|
chr7_-_27186400
|
2.237
|
NM_153715
|
HOXA10
|
homeobox A10
|
|
chr1_-_177378801
|
2.200
|
NM_001136000
NM_001168239
NM_005158
|
ABL2
|
v-abl Abelson murine leukemia viral oncogene homolog 2
|
|
chr5_+_55182963
|
2.171
|
NM_139017
|
IL31RA
|
interleukin 31 receptor A
|
|
chr9_+_130719034
|
2.145
|
|
LRRC8A
|
leucine rich repeat containing 8 family, member A
|
|
chr19_-_53305825
|
2.096
|
NM_001159322
NM_001159323
NM_003706
|
PLA2G4C
|
phospholipase A2, group IVC (cytosolic, calcium-independent)
|
|
chr22_+_44828338
|
2.094
|
|
|
|
|
chr12_+_27066749
|
2.083
|
NM_004264
|
MED21
|
mediator complex subunit 21
|
|
chr6_-_56615544
|
2.072
|
NM_001723
NM_015548
|
DST
|
dystonin
|
|
chr19_-_56946911
|
2.039
|
NM_001193306
NM_002029
|
FPR1
|
formyl peptide receptor 1
|
|
chr10_+_86174665
|
2.033
|
|
FAM190B
|
family with sequence similarity 190, member B
|
|
chr1_+_148747110
|
2.022
|
NM_004425
NM_022664
|
ECM1
|
extracellular matrix protein 1
|
|
chr7_-_87687276
|
1.974
|
|
SRI
|
sorcin
|
|
chr12_-_50871953
|
1.963
|
NM_001081492
NM_182507
|
KRT80
|
keratin 80
|
|
chr12_-_9913199
|
1.957
|
|
CLEC2B
|
C-type lectin domain family 2, member B
|
|
chr2_-_189752575
|
1.899
|
NM_000393
|
COL5A2
|
collagen, type V, alpha 2
|
|
chr7_-_13997574
|
1.887
|
NM_001163148
|
ETV1
|
ets variant 1
|
|
chr16_+_29971971
|
1.878
|
NM_000034
|
ALDOA
|
aldolase A, fructose-bisphosphate
|
|
chr16_+_29971955
|
1.877
|
|
ALDOA
|
aldolase A, fructose-bisphosphate
|
|
chr1_+_157246305
|
1.864
|
NM_005531
|
IFI16
|
interferon, gamma-inducible protein 16
|
|
chr9_-_129381086
|
1.846
|
NM_001035534
|
FAM129B
|
family with sequence similarity 129, member B
|
|
chr7_-_158304062
|
1.774
|
|
ESYT2
|
extended synaptotagmin-like protein 2
|
|
chr17_+_30594192
|
1.772
|
NM_144975
|
SLFN5
|
schlafen family member 5
|
|
chr1_+_84382529
|
1.750
|
NM_182948
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr5_+_98132898
|
1.748
|
NM_001012761
|
RGMB
|
RGM domain family, member B
|
|
chr6_+_116939500
|
1.743
|
NM_153711
|
FAM26E
|
family with sequence similarity 26, member E
|
|
chr2_-_144991386
|
1.741
|
|
ZEB2
|
zinc finger E-box binding homeobox 2
|
|
chr3_-_115825742
|
1.727
|
NM_001164347
|
ZBTB20
|
zinc finger and BTB domain containing 20
|
|
chr2_+_36777406
|
1.677
|
|
VIT
|
vitrin
|
|
chr12_+_47584159
|
1.660
|
NM_033124
|
CCDC65
|
coiled-coil domain containing 65
|
|
chr9_+_119506274
|
1.647
|
NM_138554
|
TLR4
|
toll-like receptor 4
|
|
chr6_+_121798443
|
1.645
|
NM_000165
|
GJA1
|
gap junction protein, alpha 1, 43kDa
|
|
chr9_+_74956614
|
1.645
|
|
ANXA1
|
annexin A1
|
|
chr5_+_54491702
|
1.640
|
NM_001008397
|
GPX8
|
glutathione peroxidase 8 (putative)
|
|
chr5_-_59100091
|
1.631
|
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr9_+_119506446
|
1.630
|
|
TLR4
|
toll-like receptor 4
|
|
chr9_+_119506428
|
1.628
|
|
TLR4
|
toll-like receptor 4
|
|
chr12_+_29193399
|
1.618
|
|
FAR2
|
fatty acyl CoA reductase 2
|
|
chr17_-_31219943
|
1.603
|
NM_152781
|
C17orf66
|
chromosome 17 open reading frame 66
|
|
chr3_-_130761967
|
1.584
|
|
PLXND1
|
plexin D1
|
|
chr10_+_71231693
|
1.578
|
|
COL13A1
|
collagen, type XIII, alpha 1
|
|
chr2_-_133145539
|
1.560
|
NM_001077427
|
LYPD1
|
LY6/PLAUR domain containing 1
|
|
chr5_+_96237923
|
1.535
|
|
ERAP2
|
endoplasmic reticulum aminopeptidase 2
|
|
chr8_-_13416554
|
1.509
|
NM_024767
NM_182643
|
DLC1
|
deleted in liver cancer 1
|
|
chr2_+_12775814
|
1.479
|
|
|
|
|
chr20_-_29751755
|
1.477
|
|
|
|
|
chr7_-_87687302
|
1.469
|
NM_003130
|
SRI
|
sorcin
|
|
chr1_-_242073094
|
1.467
|
NM_005465
NM_181690
|
AKT3
|
v-akt murine thymoma viral oncogene homolog 3 (protein kinase B, gamma)
|
|
chr10_+_71231649
|
1.457
|
NM_001130103
NM_080798
NM_080800
NM_080801
NM_080802
NM_080805
|
COL13A1
|
collagen, type XIII, alpha 1
|
|
chr3_-_184756101
|
1.427
|
NM_130446
|
KLHL6
|
kelch-like 6 (Drosophila)
|
|
chr2_-_215998181
|
1.416
|
|
FN1
|
fibronectin 1
|
|
chr6_+_151688327
|
1.410
|
NM_144497
|
AKAP12
|
A kinase (PRKA) anchor protein 12
|
|
chr11_-_101292462
|
1.401
|
NM_178127
|
ANGPTL5
|
angiopoietin-like 5
|
|
chr4_-_89963177
|
1.377
|
NM_001015045
|
FAM13A
|
family with sequence similarity 13, member A
|
|
chr1_+_84540921
|
1.365
|
NM_001134664
|
SAMD13
|
sterile alpha motif domain containing 13
|
|
chr2_+_102320148
|
1.356
|
NM_003856
|
IL1RL1
|
interleukin 1 receptor-like 1
|
|
chr16_+_31036474
|
1.356
|
NM_032188
NM_182958
|
MYST1
|
MYST histone acetyltransferase 1
|
|
chr17_+_73886290
|
1.354
|
NM_024419
|
PGS1
|
phosphatidylglycerophosphate synthase 1
|
|
chr9_-_116608113
|
1.350
|
NM_005118
|
TNFSF15
|
tumor necrosis factor (ligand) superfamily, member 15
|
|
chr12_-_9913724
|
1.345
|
NM_005127
|
CLEC2B
|
C-type lectin domain family 2, member B
|
|
chr3_-_71196696
|
1.332
|
|
FOXP1
|
forkhead box P1
|
|
chr12_+_55201854
|
1.325
|
NM_002898
|
RBMS2
|
RNA binding motif, single stranded interacting protein 2
|
|
chr4_+_128773529
|
1.324
|
NM_015693
|
INTU
|
inturned planar cell polarity effector homolog (Drosophila)
|
|
chr3_+_138159396
|
1.323
|
NM_144717
|
IL20RB
|
interleukin 20 receptor beta
|
|
chr3_-_170347019
|
1.312
|
NM_001105078
|
MECOM
|
MDS1 and EVI1 complex locus
|
|
chr1_+_143807748
|
1.304
|
NM_004892
|
SEC22B
|
SEC22 vesicle trafficking protein homolog B (S. cerevisiae) (gene/pseudogene)
|
|
chr1_+_84536636
|
1.293
|
NM_001010971
|
SAMD13
|
sterile alpha motif domain containing 13
|
|
chr8_+_75066117
|
1.275
|
NM_001195797
NM_015364
|
LY96
|
lymphocyte antigen 96
|
|
chr10_-_14090459
|
1.274
|
|
FRMD4A
|
FERM domain containing 4A
|
|
chr9_+_5500544
|
1.273
|
NM_025239
|
PDCD1LG2
|
programmed cell death 1 ligand 2
|
|
chr10_-_31328426
|
1.271
|
NM_001143769
|
ZNF438
|
zinc finger protein 438
|
|
chr11_+_65024889
|
1.263
|
|
MALAT1
|
metastasis associated lung adenocarcinoma transcript 1 (non-protein coding)
|
|
chr7_-_13995811
|
1.262
|
|
ETV1
|
ets variant 1
|
|
chr9_+_674193
|
1.258
|
|
|
|
|
chr2_+_170044194
|
1.250
|
NM_152384
|
BBS5
|
Bardet-Biedl syndrome 5
|
|
chr3_-_170346760
|
1.245
|
NM_001105077
NM_001163999
NM_005241
|
MECOM
|
MDS1 and EVI1 complex locus
|
|
chr8_+_104452877
|
1.244
|
|
CTHRC1
|
collagen triple helix repeat containing 1
|
|
chr5_+_40877053
|
1.244
|
NM_032587
|
CARD6
|
caspase recruitment domain family, member 6
|
|
chr20_-_14266200
|
1.235
|
NM_013281
NM_198391
|
FLRT3
|
fibronectin leucine rich transmembrane protein 3
|
|
chr1_+_84382562
|
1.224
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr11_-_8695974
|
1.220
|
|
ST5
|
suppression of tumorigenicity 5
|
|
chr9_-_116920232
|
1.213
|
NM_002160
|
TNC
|
tenascin C
|
|
chr9_-_113561613
|
1.213
|
NM_001080551
|
C9orf84
|
chromosome 9 open reading frame 84
|
|
chr15_+_37330176
|
1.212
|
NM_207445
|
C15orf54
|
chromosome 15 open reading frame 54
|
|
chr22_-_43987978
|
1.211
|
|
KIAA0930
|
KIAA0930
|
|
chrX_-_19598912
|
1.207
|
NM_001184960
|
SH3KBP1
|
SH3-domain kinase binding protein 1
|
|
chrX_+_15428820
|
1.187
|
NM_203281
|
BMX
|
BMX non-receptor tyrosine kinase
|
|
chr7_+_28692122
|
1.175
|
NM_001011666
|
CREB5
|
cAMP responsive element binding protein 5
|
|
chr14_+_50096603
|
1.167
|
|
ATL1
|
atlastin GTPase 1
|
|
chr4_-_70861301
|
1.163
|
NM_001891
|
CSN2
|
casein beta
|
|
chr6_-_134540666
|
1.161
|
NM_001143677
|
SGK1
|
serum/glucocorticoid regulated kinase 1
|
|
chr12_+_10256682
|
1.155
|
NM_031412
|
GABARAPL1
|
GABA(A) receptor-associated protein like 1
|
|
chr6_-_112682401
|
1.153
|
|
LAMA4
|
laminin, alpha 4
|
|
chr16_+_27248806
|
1.148
|
|
IL4R
|
interleukin 4 receptor
|
|
chr4_-_186693574
|
1.145
|
NM_001114107
NM_014476
|
PDLIM3
|
PDZ and LIM domain 3
|
|
chr7_-_112513379
|
1.126
|
NM_001146266
NM_001146267
|
GPR85
|
G protein-coupled receptor 85
|
|
chr8_-_102029997
|
1.121
|
|
YWHAZ
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide
|
|
chr9_-_16717887
|
1.115
|
|
BNC2
|
basonuclin 2
|
|
chr18_-_21184824
|
1.112
|
|
|
|
|
chr21_+_42492867
|
1.111
|
NM_207627
NM_207628
|
ABCG1
|
ATP-binding cassette, sub-family G (WHITE), member 1
|
|
chr11_+_113436438
|
1.111
|
NM_001018011
|
ZBTB16
|
zinc finger and BTB domain containing 16
|
|
chr1_+_242691295
|
1.102
|
NM_001130957
NM_173807
|
C1orf101
|
chromosome 1 open reading frame 101
|
|
chr5_+_159546951
|
1.100
|
NM_001130958
|
FABP6
|
fatty acid binding protein 6, ileal
|
|
chr17_+_55098280
|
1.088
|
|
CLTC
|
clathrin, heavy chain (Hc)
|
|
chr8_+_104452961
|
1.086
|
NM_138455
|
CTHRC1
|
collagen triple helix repeat containing 1
|
|
chr12_+_118100977
|
1.083
|
NM_014365
|
HSPB8
|
heat shock 22kDa protein 8
|
|
chr1_-_67366807
|
1.080
|
NM_001013674
|
C1orf141
|
chromosome 1 open reading frame 141
|
|
chr6_-_112682459
|
1.080
|
NM_001105206
NM_001105207
NM_001105208
NM_001105209
NM_002290
|
LAMA4
|
laminin, alpha 4
|
|
chr13_+_96726197
|
1.071
|
|
MBNL2
|
muscleblind-like 2 (Drosophila)
|
|
chr1_-_85515174
|
1.066
|
NM_003921
|
BCL10
|
B-cell CLL/lymphoma 10
|
|
chr11_+_57547928
|
1.055
|
NM_001005212
|
OR9Q1
|
olfactory receptor, family 9, subfamily Q, member 1
|
|
chr9_+_123123364
|
1.036
|
|
GSN
|
gelsolin
|
|
chr1_-_151866147
|
1.034
|
NM_001024213
|
S100A13
|
S100 calcium binding protein A13
|
|
chr3_-_188946168
|
1.027
|
NM_001706
|
BCL6
|
B-cell CLL/lymphoma 6
|
|
chr2_+_36777336
|
1.027
|
NM_001177969
NM_001177970
NM_001177971
NM_001177972
NM_053276
|
VIT
|
vitrin
|
|
chr1_+_156229686
|
1.023
|
NM_018240
|
KIRREL
|
kin of IRRE like (Drosophila)
|
|
chr7_+_150442652
|
1.022
|
|
AGAP3
|
ArfGAP with GTPase domain, ankyrin repeat and PH domain 3
|
|
chr4_+_75393053
|
1.020
|
NM_001013442
|
EPGN
|
epithelial mitogen homolog (mouse)
|
|
chr1_-_171441234
|
1.020
|
|
TNFSF4
|
tumor necrosis factor (ligand) superfamily, member 4
|
|
chr6_-_112682573
|
1.013
|
|
LAMA4
|
laminin, alpha 4
|
|
chr9_-_4290028
|
1.012
|
NM_001042413
|
GLIS3
|
GLIS family zinc finger 3
|
|
chr1_+_99884018
|
1.002
|
NM_017734
|
PALMD
|
palmdelphin
|
|
chr7_+_120415958
|
0.988
|
NM_024913
|
C7orf58
|
chromosome 7 open reading frame 58
|
|
chr6_-_128883125
|
0.980
|
NM_002844
NM_001135648
|
PTPRK
|
protein tyrosine phosphatase, receptor type, K
|
|
chr15_+_73803351
|
0.969
|
NM_175881
|
ODF3L1
|
outer dense fiber of sperm tails 3-like 1
|
|
chr10_+_53744043
|
0.965
|
NM_012242
|
DKK1
|
dickkopf homolog 1 (Xenopus laevis)
|
|
chr9_-_4289915
|
0.961
|
|
GLIS3
|
GLIS family zinc finger 3
|
|
chr9_+_12764988
|
0.957
|
NM_203403
|
C9orf150
|
chromosome 9 open reading frame 150
|
|
chr5_+_34723420
|
0.940
|
NM_001145523
|
RAI14
|
retinoic acid induced 14
|
|
chr12_-_69317441
|
0.939
|
NM_001109754
|
PTPRB
|
protein tyrosine phosphatase, receptor type, B
|
|
chr11_+_59279481
|
0.938
|
|
STX3
|
syntaxin 3
|
|
chr3_-_11598829
|
0.936
|
NM_001128220
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr9_-_109291575
|
0.936
|
|
KLF4
|
Kruppel-like factor 4 (gut)
|
|
chr4_+_74825138
|
0.931
|
NM_000584
|
IL8
|
interleukin 8
|
|
chrX_-_23955223
|
0.923
|
NM_030624
|
KLHL15
|
kelch-like 15 (Drosophila)
|
|
chr2_-_40593074
|
0.921
|
NM_001112802
|
SLC8A1
|
solute carrier family 8 (sodium/calcium exchanger), member 1
|
|
chr19_-_42393225
|
0.918
|
NM_152279
|
ZNF585B
|
zinc finger protein 585B
|
|
chr14_+_50096492
|
0.916
|
NM_015915
NM_181598
|
ATL1
|
atlastin GTPase 1
|
|
chr2_-_188021117
|
0.900
|
NM_005795
|
CALCRL
|
calcitonin receptor-like
|
|
chr3_+_137224266
|
0.892
|
NM_181897
|
PPP2R3A
|
protein phosphatase 2, regulatory subunit B'', alpha
|
|
chr4_+_124537334
|
0.892
|
NM_199327
|
SPRY1
|
sprouty homolog 1, antagonist of FGF signaling (Drosophila)
|
|
chr12_-_14929991
|
0.887
|
NM_000900
NM_001190839
|
MGP
|
matrix Gla protein
|
|
chr2_-_207738858
|
0.883
|
NM_003709
|
KLF7
|
Kruppel-like factor 7 (ubiquitous)
|
|
chr1_-_110735158
|
0.881
|
NM_004696
|
SLC16A4
|
solute carrier family 16, member 4 (monocarboxylic acid transporter 5)
|
|
chr19_-_51319769
|
0.881
|
NM_207393
|
IGFL3
|
IGF-like family member 3
|
|
chr10_+_118177371
|
0.880
|
NM_001011709
|
PNLIPRP3
|
pancreatic lipase-related protein 3
|
|
chr22_+_43527101
|
0.880
|
NM_001017526
NM_001198726
NM_181335
|
ARHGAP8
|
Rho GTPase activating protein 8
|
|
chr11_-_121492010
|
0.870
|
NM_001001786
|
BLID
|
BH3-like motif containing, cell death inducer
|
|
chr1_+_84382575
|
0.869
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr1_+_218768146
|
0.867
|
|
MARK1
|
MAP/microtubule affinity-regulating kinase 1
|
|
chr5_-_119768430
|
0.863
|
|
|
|
|
chr11_+_59279377
|
0.861
|
|
STX3
|
syntaxin 3
|
|
chr18_+_7936885
|
0.860
|
|
PTPRM
|
protein tyrosine phosphatase, receptor type, M
|
|
chr16_+_28993770
|
0.859
|
|
RRN3P2
|
RNA polymerase I transcription factor homolog (S. cerevisiae) pseudogene 2
|
|
chr3_-_102194938
|
0.856
|
NM_015429
|
ABI3BP
|
ABI family, member 3 (NESH) binding protein
|
|
chr11_+_111552298
|
0.852
|
NM_001037290
|
BCO2
|
beta-carotene oxygenase 2
|
|
chr10_+_94584449
|
0.848
|
NM_001013848
|
EXOC6
|
exocyst complex component 6
|
|
chrX_+_100361588
|
0.844
|
NM_001171184
NM_001939
|
DRP2
|
dystrophin related protein 2
|
|
chr1_-_151866367
|
0.844
|
NM_001024212
|
S100A13
|
S100 calcium binding protein A13
|
|
chr7_-_27125738
|
0.843
|
NM_030661
|
HOXA3
|
homeobox A3
|
|
chr9_+_138677197
|
0.841
|
NM_201446
NM_016215
|
EGFL7
|
EGF-like-domain, multiple 7
|
|
chr2_-_225519974
|
0.840
|
|
DOCK10
|
dedicator of cytokinesis 10
|
|
chr11_+_111551417
|
0.840
|
NM_031938
|
BCO2
|
beta-carotene oxygenase 2
|
|
chr21_-_35174742
|
0.839
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr8_+_67201684
|
0.838
|
NM_033058
NM_184085
NM_184086
NM_184087
|
TRIM55
|
tripartite motif containing 55
|
|
chr9_-_25668855
|
0.838
|
NM_001004125
|
TUSC1
|
tumor suppressor candidate 1
|
|
chr3_+_87359088
|
0.838
|
NM_014043
|
CHMP2B
|
chromatin modifying protein 2B
|